Unexpected effects on embryonic development are caused by ancient, dormant sequences in the genome. This is because retroviral sequences that are mostly “harmless” yet undead are present in the mammalian genome.
Recently, some of these retroviral gene fragments’ potential effects on embryonic cells were discovered by an international research team. Unexpectedly, an imbalance in the cell is caused by copies of the genetic material rather than the viral proteins.
“We found that the messenger copies of some of the viral genes, the RNA, have an important impact on embryonic cells. The viral sequences seem to remember their original mission of hijacking the molecular machinery that ensures the flow of information from DNA to RNA to protein. Interestingly, the messenger RNA itself seems to be responsible,”
research leader Denes Hnisz, Max Planck Institute for Molecular Genetics (MPIMG) in Berlin, said.
Ancient Retroviral Sequences
Numerous viruses have integrated themselves into our genome over the course of thousands of years of evolution. Ancient retroviral sequences make up a startling 10% of mammalian genomes.
These no longer appear to be a threat because the majority of them have mutated beyond recognition. Furthermore, the cell has epigenetically silenced these genes.
However, if the viral remains are not silenced, they will rise from their graves, causing chaos in the cell.
The RNA of the resurrected viruses exerts attractive forces on the enzymes that read the information from the DNA, according to Hnisz’ team and collaborators. The embryonic cell’s tasks, such as reading important embryonic genes, are neglected, resulting in a fatal imbalance.
For instance, certain forms of cancer and neurological disorders exhibit this unleashed state.
Genetic Alteration
Viruses are shrewdly designed genetic informational fragments. Some of them embed themselves into their hosts’ genomes and stay there. Endogenous Retroviruses (ERVs) have multiplied to thousands of copies, frequently in swarms of hundreds of repetitive copies, throughout mammalian genomes.
“As retroviruses jump from one section of DNA to the next during their life cycle, they can alter genes and even recombine them. This makes them an important tool for evolution to create new genes,”
said Henri Niskanen, one of the scientists involved in the study.
Uncontrolled gene modification, however, is not good for an individual organism, particularly during the development of the embryo. Because of this, the cell will recognize endogenous retrovirus sequences and send specialized repressive machinery to their locations in order to silence them. Additionally, at these locations, the chromosome is compacting.
Missing Trim28
But what happens if you disable these safety features? The research team was interested in learning what occurs as soon as the ERV zombies are no longer under control. To achieve this, they took Trim28, a protein that silences viral remnants, out of mouse embryonic stem cells, and observed the immediate effects.
When Trim28 was removed, the cell read more ERV genes, producing RNA copies with the assistance of the RNA polymerase enzyme. However, the polymerase unexpectedly disappeared from stem cell genes that are critical for stem cell potency.
Each cell has a finite supply of polymerase enzymes and other required factors. When too many genes are transcribed at once, they compete for limited resources. In an experiment, repeats of ERV sequences competed against stem cell genes.
“We see that ERV repeats have a slightly higher affinity. They draw the machinery away from embryonic genes, creating an imbalance,”
said Christina Riemenschneider, another of the team’s researchers.
Stem Cell Impact
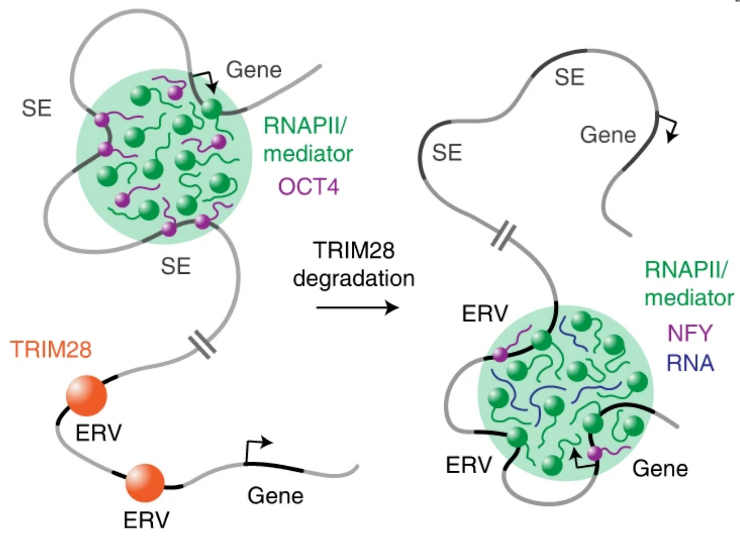
RNA polymerase and other necessary factors that selectively dock onto genes frequently assemble into droplets that float around in the cell nucleus, similar to oil droplets in salad dressing. These “condensates” contain many of the molecules required for gene reading and are especially drawn to specific DNA segments that control the most important genes in a cell.
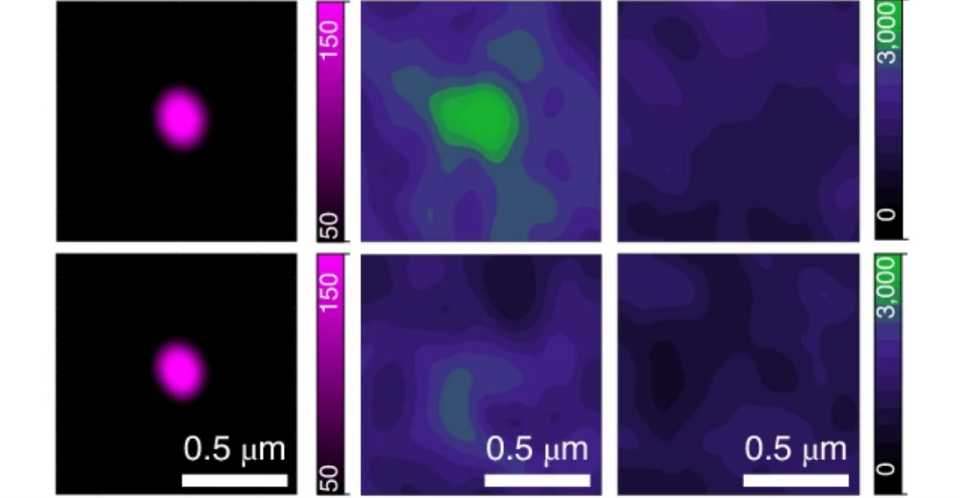
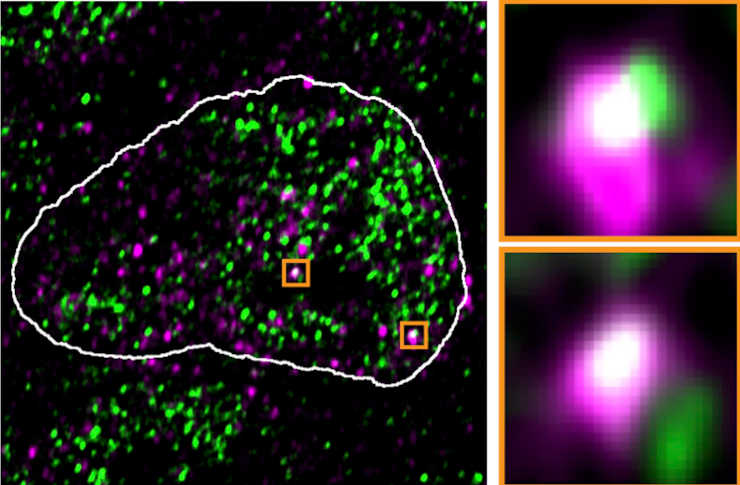
The ERV genes, or more precisely the RNA molecules produced by these genes, appeared to take over the condensates. They were frequently found in the same locations as the reactivated ERV genes in high-resolution microscopic images. The droplets returned to their original location after the viral RNA was removed from the cells.
The virus-like RNA’s effects were not limited to the molecular level. Working in early mouse embryos, the researchers discovered that shifting the condensates toward the ERVs had a negative impact on development.
Because necessary genes were no longer active, stem cells, for example, lost their characteristic ability to develop into any other cell.
“It’s quite remarkable that non-coding, non-functional genes have such a profound effect via RNA. You might imagine DNA damage or viral particles when thinking of retroviruses that integrate into the genome, but that is not the case in this instance,”
said scientist Abhishek Sampath Kumar. The team’s discovery sheds new light on endogenous retrovirus research.
“Hijacking of transcriptional condensates by the ERVs and their RNA is an important mechanistic finding that should be taken into account in future studies of transposable elements and their epigenetic regulators. This could be an additional route ERVs used to contribute to evolutionary innovation,”
said researcher Vahid Asimi.
ERV reactivation is clearly linked to pathologies ranging from obesity to various cancers to neurological diseases like amyotrophic lateral sclerosis and schizophrenia. Denes Hnisz, the group’s leader, hopes that the research will aid in determining the molecular causes of these diseases.
Reference: Asimi, V., Sampath Kumar, A., Niskanen, H. et al. Hijacking of transcriptional condensates by endogenous retroviruses. Nat Genet 54, 1238–1247 (2022).